Exploring Protein Interactions In Budding Yeast
25 Sep 2014Prologue
I've been away longer than I'd like to have been. Generally I strive for a once a week clip in posting. But life has pulled me in different directions lately and my mind has been on other matters. Lately, my wife and I have been anticipating a new addition to our family early next year, a baby girl.
As I've played around with several graph datasets (which will be featured in future posts) I settled on a dataset from biology, since my mind seems to have been preoccupied with that topic. I thought this could be a catalyst to bring my mind back to this project. What I quickly learned is that I am not a biology expert and exploring this dataset required my learning new vocabulary and tools. This took extra time and effort, but was also very enjoyable as I'll describe below.
Intro
The dataset I'm starting with for this is the Pajek Protein-protein interaction network in budding yeast. The network has 2361 proteins and models 6646 protein interactions. I filtered it to only proteins that have more than one interaction to remove proteins that are generally less impactful on the network. This filtered it down to 1528 proteins and 5913 protein interactions. I loaded it up into Gephi and did a typical analysis. See my post on analyzing your Facebook friend network for a step-by-step set of instructions on how to do this. For my rank sorting I chose to use a betweeness centrality algorithm. This measures how often a node (or in this case a protein) appears on pathways between other nodes in the network. In a biological sense, if there are processes that involve multiple interactions between different proteins, then the proteins with high betweeness centrality scores are more likely to play a role in these processes.
I used an Open Ord layout, which is optimized for larger networks. The end result looks something like this:

Here's an SVG version.
A Note on Systems Biology
The field of systems biology has emerged recently as our tools for modeling complex systems have improved. This is particularly relevant for graph analysis as graph tools are designed specifically to understand the interactions and interrelationships between actors or elements in a system. It reminds me of Warren Weaver's foresight back in 1948. In an essay titled "Science and Complexity" Warren Weaver points to how improvements in computational tools would allow new fields of study and discipline to emerge in the study of systems:
The combination of flexibility, capacity, and speed makes it seem like that [computers] will have a tremendous impact on science. That will make it possible to deal with problems which previously were too complicated, and, more importantly, they will justify and inspire the development of new methods of analysis applicable to these new problems of organized complexity.
...these new ways of working, effectively instrumented by huge computers, will contribute greatly to the advance which the next half century will surely achieve in handling the complex, but essentially organic, problems of the biological and social sciences.
Warren Weaver's essay has been a great inspiration for me in doing this blog. And with regards to an emerging field of systems biology made possible through advances in computational modeling tools and techniques, Warren Weaver's foresight has proven to be quite prophetic.
Key Proteins
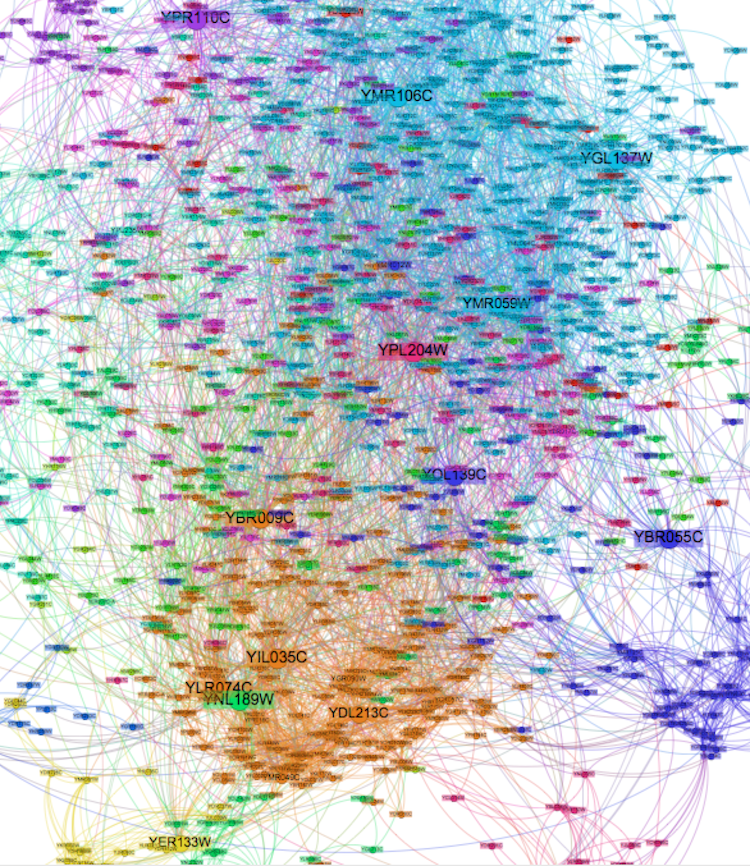

After doing a centrality scoring, several proteins emerged as playing frequent roles in protein interactions for this population (in order of betweeness score):
At this point I was probably as confused as you might be. These labels mean nothing to me. Fortunately, the name is all that is required to look these proteins up in various biology databases such as the one used in the links above. I found that the Saccharomyces Genome Database provided more than enough information on each of these proteins.
Visualization
Even with a genome database of information about each protein, I was still at a loss of what to make of this list. After some exploration, I found that I could visualize known protein structures with sequence similarity to the ones listed above. These are known as biological homologs. Furthermore, I wanted to (where possible) select homologs that are known to appear in the human body. I used the tool Jmol to do this visualization.
Here's a close up inside one of these proteins when visualized:
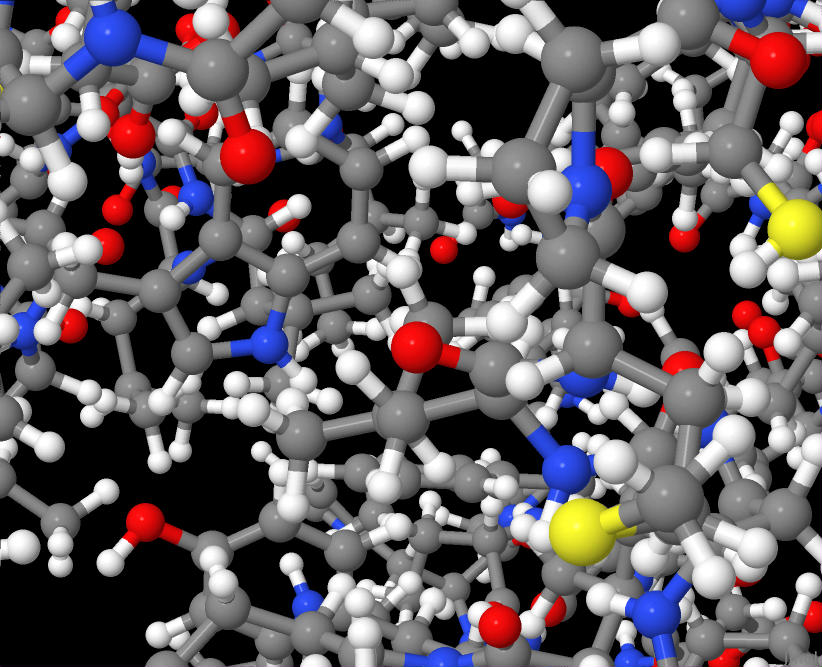
Notice that molecules themselves are graphs with atoms as nodes and bonds as edges. In fact, networks are used to model chemical and atomic processes themselves as well as other complex systems as is the case in Bose-Einstein condensation and biological networks in general.
There is probably much, much more that could be analyzed from this network and scoring of proteins, but I just wanted to simply show a visualization of the proteins that scored high themselves and highlight the universality of networks as their models emerge out of both the microscopic and the macroscopic levels.
YNL189W 2Z6H
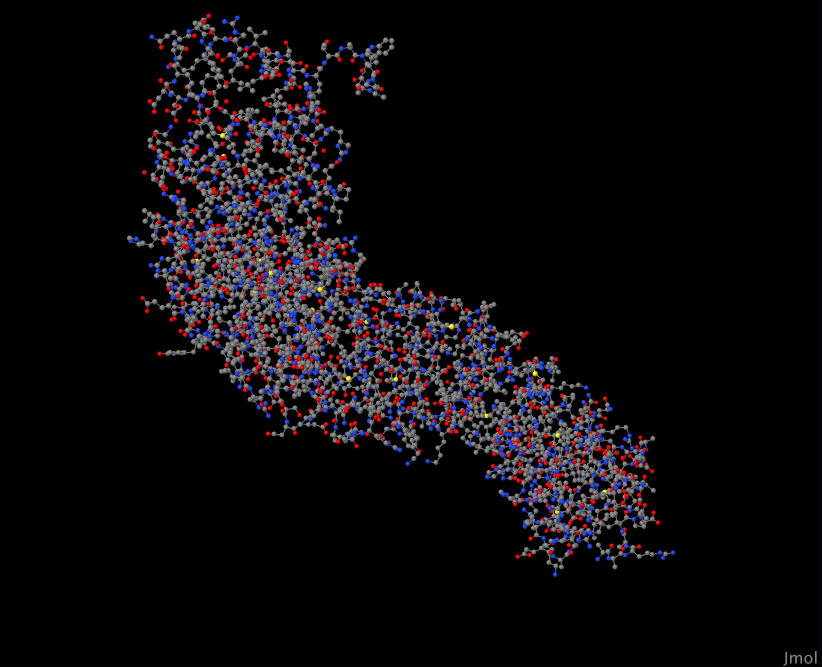
YMR106C
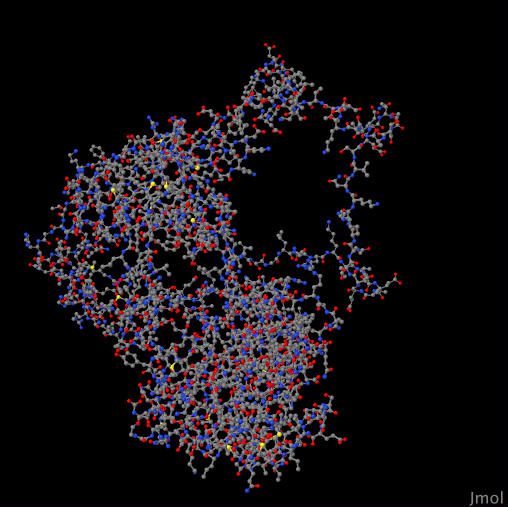

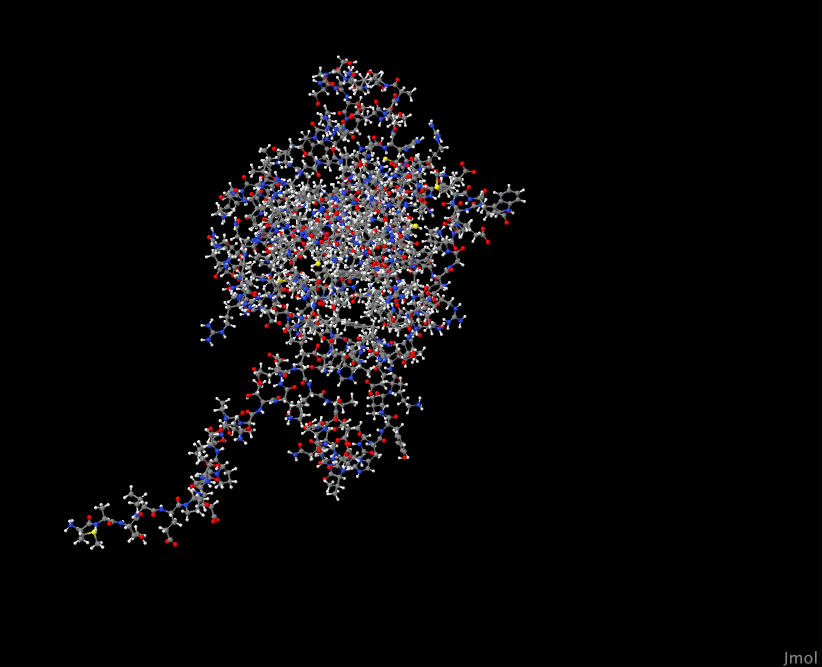
YPL204W 4HOK
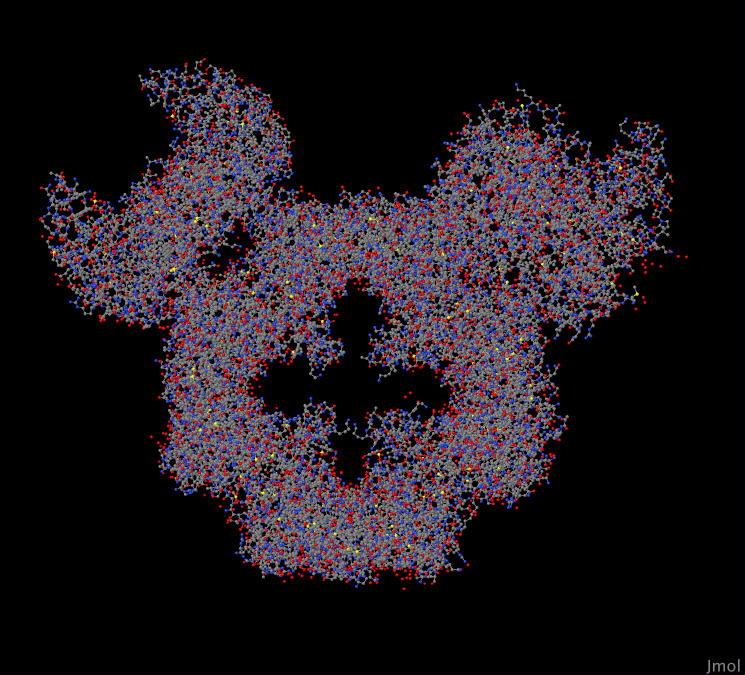

YPR110C 3J0K
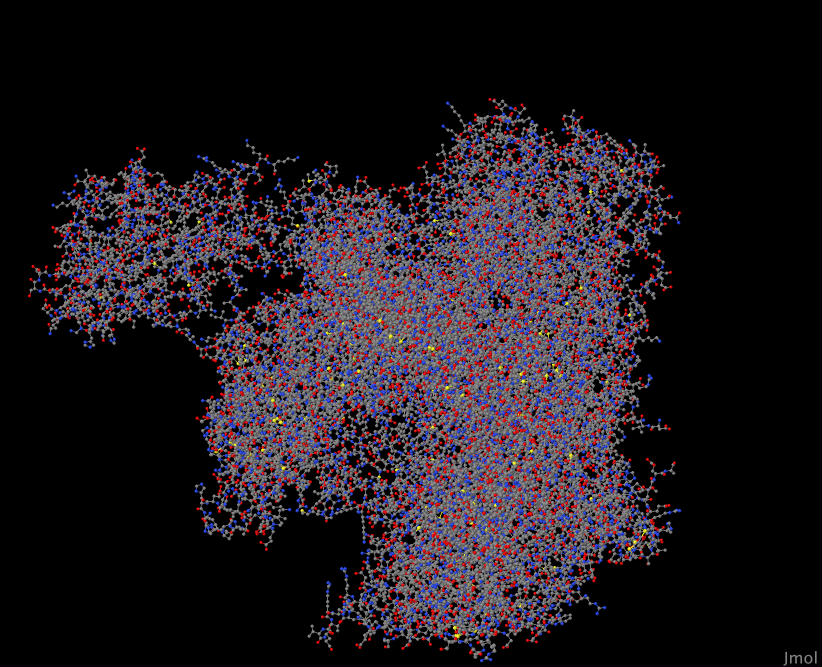

YGL137W 3EMH
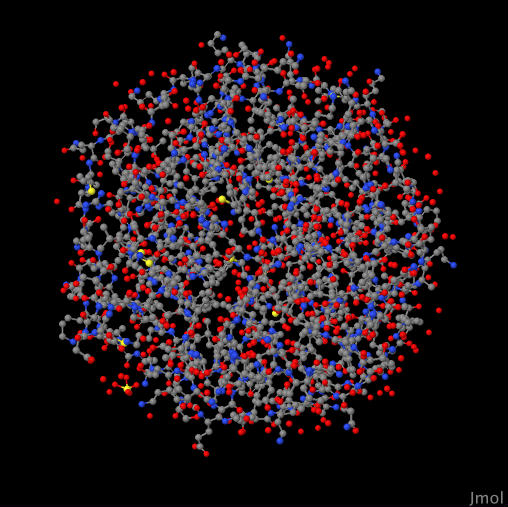

YBR009C 3AZI
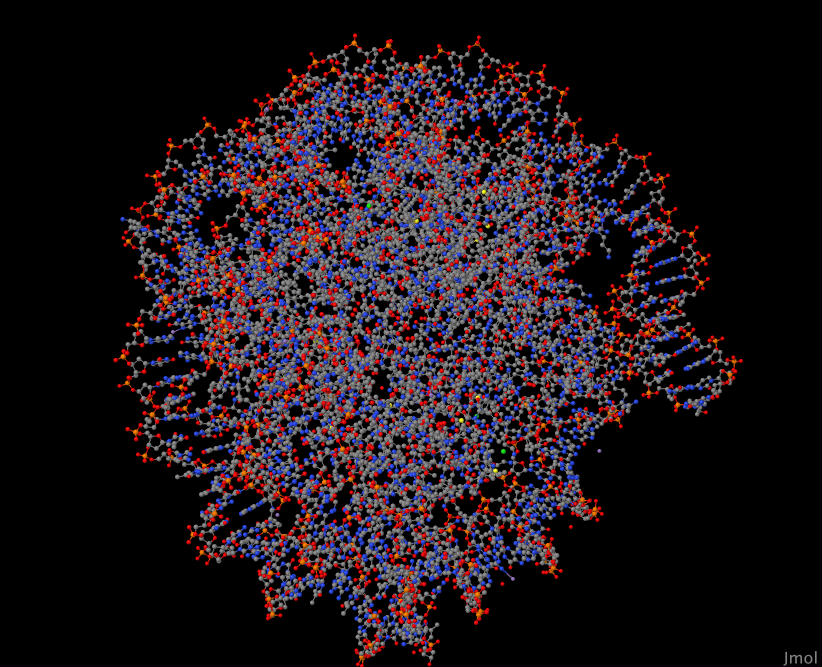

YIL035C 3WAR
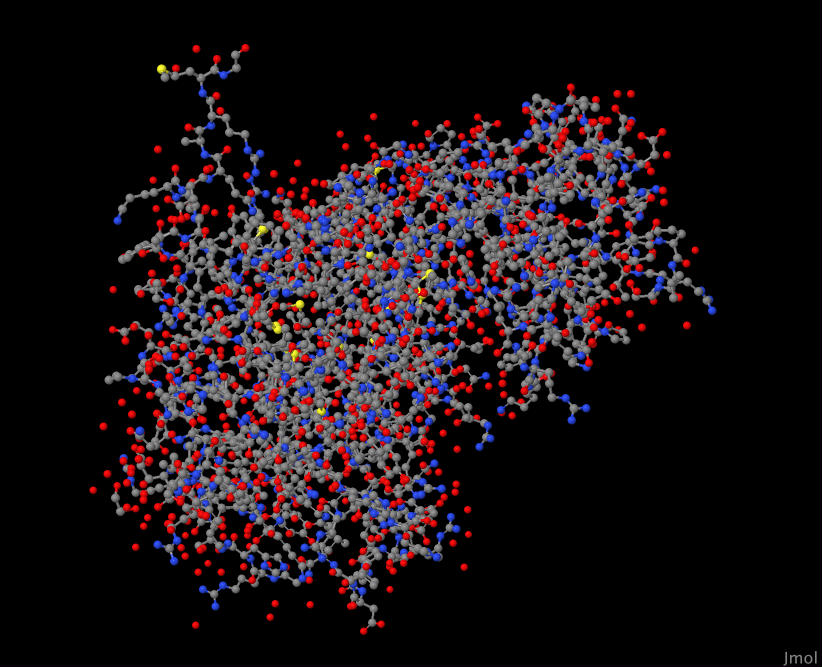

YLR074C 1ZR9
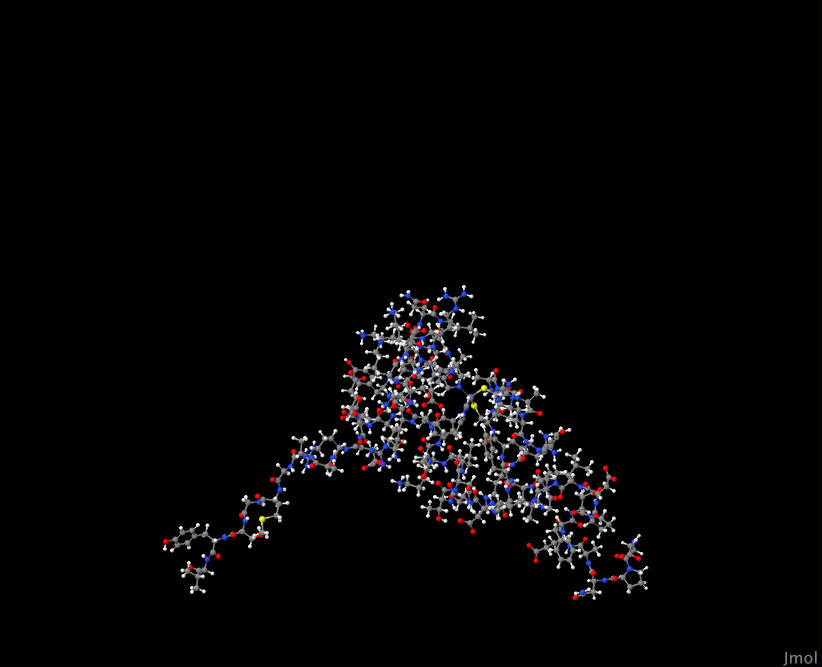

YOL139C 2GPQ
 AllThingsGraphed.com
AllThingsGraphed.com